Team:NC School of Sci Math/Lab Notebook
From 2013hs.igem.org
Home Team Project Details Lab Notebook Results Human Impact Biosafety Acknowledgments Official Team Profile
Planning and Development
01/11/2013
Brainstorming for project ideas- LDL-cholesterol biosensor
- Ocean salinity regulator (especially for coral reefs)
- Water contaminant multibiosensor
More research should be done
- Find related projects that have been completed
- Obstacles?
- BioBricks?
- Presentations in 3-4 weeks
01/18/2013
Discussions of each project- LDL-cholesterol biosensor
- Could be used as a blood test
- Blood sugar monitor-like device
- Hard to measure any color change in blood
- Ocean salinity regulator (especially for coral reefs)
- Need a lot of bacteria
- Environmental concerns
- Bacteria can do well in coral reefs
- Water contaminant multibiosensor
- Several Biobrick promoters for common contaminants
- Usage in sewer tanks
- Many different colors of fluorescent proteins
- Could incorporate newer technology for better color-sensing
- Will look up data on promoter/FP expression
02/01/2013
Presentations
- All ideas sound promising
- Meet next week to decide which idea to pursue
02/08/2013
Decision on project- LDL-cholesterol idea won't work
- Costly and hard to detect FPs in blood
- Multibiosensor is appealing
- Many applications
- Expandable with many different FPs
- Could bring in new technology
- Ocean salinity regulator is difficult because of the ocean
- Environmental factors and unknown variables
Multibiosensor chosen
02/15/2013
A few members met with Mr. Jon Davis to discuss potential technology
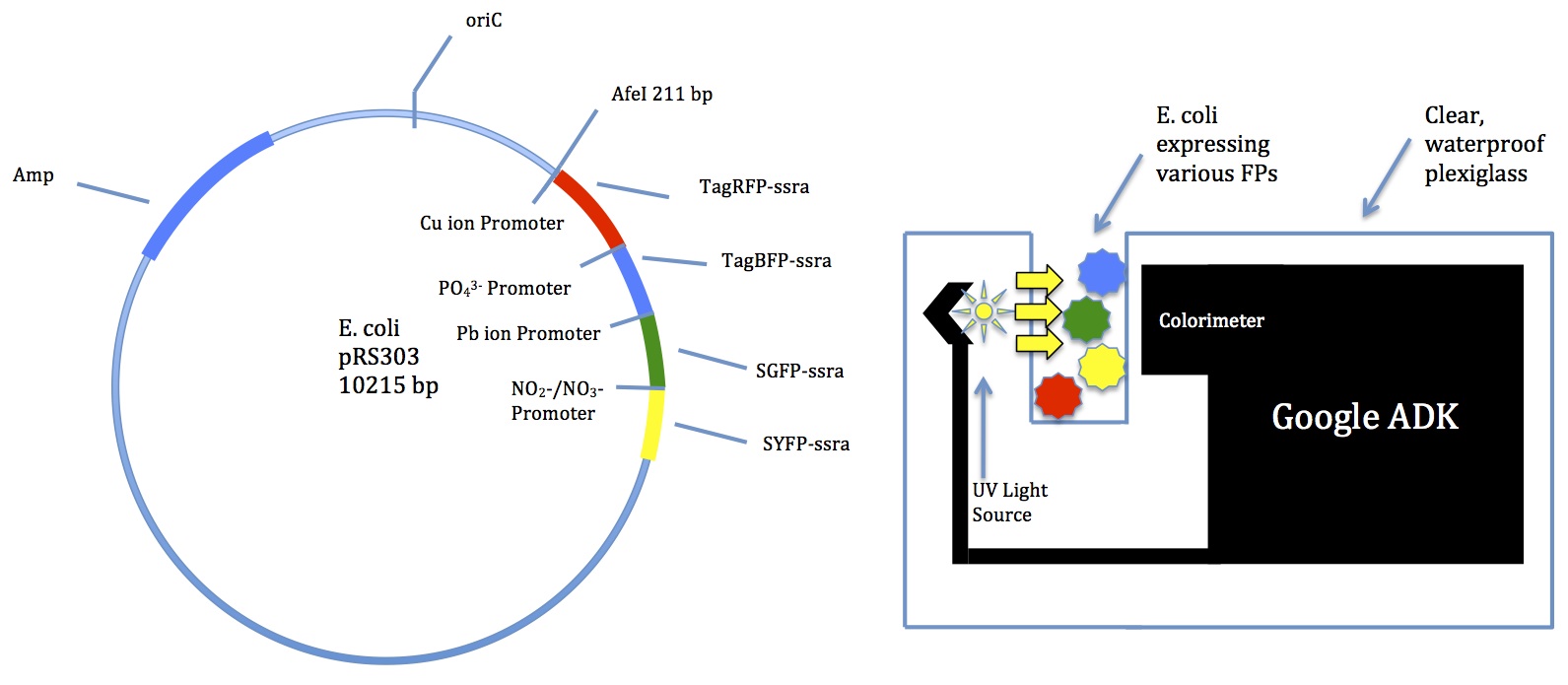
- Suggested a Google device called ADK
- Comes with a colorimeter
- Can be programmed for many functions
- School has a few we could try out
02/22/2013
Goals for the immediate future
- Find Biobricks
- Set up schedule for lab work
- Further research into possible uses
- Preliminary lab experiments
Project Proposal
03/01/2013
Discussion of upcoming project proposal:
- Introduction, Solution Statement, and Methods
- Need to find out some background information
- Should make use of Tinkercell modeling
03/04/2013
Assignments of roles for the upcoming project proposal:
- Madeline will investigate background information
- Danny will work on Tinkercell models
- Jack will write the solution statement and methods sections
- Synthesize our writing on a Google Doc
- Try to have individual parts done in about 2 weeks
03/18/2013
The following models have been created:
- Gelelectrophoresis of the digested biobrick parts in order to extract the GFP and LacZ backbone as well as precA
Protocol:
In the meantime, the restriction digests were mixed with 6 µl of 6 x Loading Dye (fermentas). The whole restriction digest was loaded onto a 1 % agarose gel. Therefore, the agarose gel was prepared by adding 2 g of agarose to 200 ml of 1x TBE buffer and heated up in a microwave for 2 min/600 W. Thereby, the agarose was melted. Afterwards, the gel was stirred on a magnetic stirring device until the solution reached ~ 60°C and then the gel was poured. 10 µl of ethidium bromide were subsequently added by our supervisor Katharina Genreith (as EtBr is very toxic, we preferred our supervisor to do the handling of that when running a gel for the first time).
The samples were then loaded on the gel together with 1kb plus loading ladder (fermentas) and run for 1 h @ 100V. The expected bands can be seen on fig. 1.

Fig. 1: Schema of the gel we expect based on the length(picture constructed using the WinSerial Cloner software)

Fig. 2: Picture taken of the gel. All expected bands are visible
- Gel extraction of the indicated band (backbones for LacZ and GFP as well as precA insert) was done using a Qiagen gel extraction kit.
Protocol:
The bands were cut of the gel under UV light (caution: wear goggles in order to protect your eyes!) using a scalpel and the extracted band was put into a 2 ml eppi cup.

Fig. 3: Picture taken of the cut gel. This picture was taken after the required bands were excised for gel extraction
Afterwards, 600 µl of buffer QG were added to each gel band and the mixture was incubated on a thermomixer device at 50 °C and 700 rpm shaking for 20 min. Afterwards, 200 µl of Isopropanol were added to the mix containing precA (as precA is pretty short compared to the others, isopropanol is required). Afterwards, the whole reaction mix was loaded onto a gel extraction column and centrifuged for 1 min/max speed; flow-through was discarded. The column was subsequently washed with 500 µl of buffer QG and then 750 µl buffer PE, always centrifuging the column at max speed for 1 min at each washing step and discarding the flow-through. Finally, DNA was eluted in 20 µl of water.
- Concentration measurement of the pre-cut parts using Nanodrop
| Part | DNA concentration |
| GFP | 12.5 ng/µl |
| LacZ | 15.11 ng/µl |
| RecA | 6.0 ng/µl |
03/13/2012
- Ligation reactions were set up using 3-6 fold access of insert (precA) and 50 ng of each backbone (roughly).
- precA-GFP: 4 µl (~50 ng) GFP-backbone + 1,5 µl (~9 ng) RecA + 2 µl T4 DNA ligase buffer + 11,5 µl water + 1 µl T4 ligase
- precA-LacZ: 3 µl (~50 ng) LacZ-backbone + 1,5 µl (~9 ng) RecA + 2 µl T4 DNA ligase buffer + 12 µl water + 1 µl T4 ligase
- psulA-GFP: 4 µl (~50 ng) GFP-backbone + 1 µl annealed SulA-oligos + 2 µl buffer + 12 µl water + 1 µl T4 ligase
- psulA-LacZ: 3 µl (~50 ng) LacZ-backbone + 1 µl annealed SulA-oligos + 2 µl buffer + 13 µl water + 1 µl T4 ligase
- Incubation for 50 min at room temperature
- Heat inactivation of the T4 ligase at 70 °C for 5 min
- Afterwards, 10 µl of the ligation mix (chilled) were used for transforming 50 µl of E. coli Top10 chemocompetent cells.
03/14/2012
- Screening of the four different biobrick clonings done by colony-PCR.
Protocol:
A reaction mix was prepared containing 0.2 µl of each screening primer used for the screening, 9.6 µl of water and 10 µl of 2x PCR mastermix (fermentas). Finally, a colony was picked from a plate using a sterile pipette tip and was dipped into the PCR mix a few times. 7-8 clones were screened for each different cloning setup as follows:
- psulA-GFP and psulA-LacZ: primers psulA_fw and VR (standard sequencing primer) were used. Only in case psulA was successfully cloned into the backbone containing the reporter gene, we would get a PCR product. GFP would give a product of roughly 1000 bp, LacZ one of roughly 3500 bp.
- precA-GFP and precA-LacZ: primers VF2 and VR (standard sequencing primers) were used. We compared the product size of the different clones to the product size from the original vectors from the registry (only containing recA or LacZ). In case the cloning was successful, we should see a 200 bp shift in product size, which can be detected by gel electrophoresis.
- psulA-GFP and psulA-LacZ: primers psulA_fw and VR (standard sequencing primer) were used. Only in case psulA was successfully cloned into the backbone containing the reporter gene, we would get a PCR product. GFP would give a product of roughly 1000 bp, LacZ one of roughly 3500 bp.
The PCR program was done as follows:
94°C/3min </div>
 "
"